What's new
- 2016.01.29
"Japanese Quail (Coturnix japonica) Genome Sequencing Project" website was released.
Coturnix japonica
The Japanese quail (Coturnix japonica, Phasianidae, Galliformes), also known as Coturnix quail, is a species of Old World quail found in East Asia.
First considered a subspecies of the Common quail, it was distinguished as its own species in 1983. The Japanese quail and chicken are estimated to have diverged 35million years ago (MYA) by molecular phylogenetic analysis (van Tuinen et al. 2004). The Japanese quail has played an active role in the lives of humanity since the 12th century, and continues to play major roles in industry and scientific research. This migratory bird, with its small body size (100~300 g), short generation interval, and high resistance to disease, is an excellent model animal for a wide range of scientific research. In addition, this species is economically important as an egg- and meat-producing agricultural animal that is reared in many countries worldwide. Although the Japanese quail has several advantages as a laboratory animal for biological and biomedical investigations and as a pilot animal for poultry science, the development of quail genome information has been still insufficient. This prevents the practical use of this animal species. We here open the genome browser of the Japanese quail as a web-based tool, which has been built using our most recently updated data of the quail genome assembly.
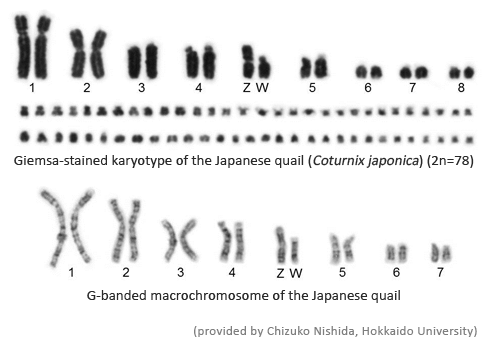
【Karyotype】【Resource】【Genome sequence information】 【Contribution】 【Members of consortium】 【References】
| Library (insert size) | # of libraries | Read length (bp) | Raw | Filter | Sequencing Platforms | ||||
|---|---|---|---|---|---|---|---|---|---|
| Number of reads (million) | Number of bases (giga) | Coverage (x) | Number of reads (million) | Number of bases (giga) | Coverage (x) | ||||
| Pair-end (300 bp) | 2 | 100 | 97.4 | 9.7 | 10.4 | 91.2 | 8.4 | 9.0 | Illumina HiSeq 2000 |
| Pair-end (500 bp) | 2 | 100 | 60.4 | 6.0 | 6.4 | 58.6 | 5.5 | 5.9 | Illumina HiSeq 2000 |
| Pair-end (300 bp) | 1 | 101 | 223.9 | 22.6 | 24.1 | 206.2 | 19.5 | 20.8 | Illumina HiSeq 2000 |
| Pair-end (500 bp) | 1 | 101 | 221.4 | 22.4 | 23.8 | 202.2 | 19.2 | 20.5 | Illumina HiSeq 2000 |
| Pair-end (300 bp) | 1 | 102 | 136.3 | 13.9 | 14.8 | 124.4 | 11.7 | 12.5 | Illumina HiSeq 2000 |
| Pair-end (500 bp) | 1 | 102 | 193.7 | 19.8 | 21.1 | 178.7 | 16.9 | 18.1 | Illumina HiSeq 2000 |
| Pair-end PCR-free (300 bp) | 1 | 100 | 865.3 | 86.5 | 92.3 | 809.7 | 73.1 | 77.9 | Illumina HiSeq 2000 |
| Pair-end PCR-free (500 bp) | 1 | 100 | 538.3 | 53.8 | 57.4 | 473.2 | 38.1 | 40.6 | Illumina HiSeq 2000 |
| Mate-pair (3 kbp) | 2 | 36 | 101.2 | 3.6 | 3.9 | 90.8 | 3.3 | 3.5 | Illumina GAII |
| Mate-pair (5 kbp) | 2 | 36 | 45.4 | 1.6 | 1.7 | 28.6 | 1.0 | 1.1 | Illumina GAII |
| Mate-pair (10 kbp) | 1 | 100 | 408.4 | 40.8 | 43.6 | 385.7 | 35.7 | 38.1 | Illumina HiSeq 2000 |
| 10X Genomics linked-reads | 1 | 100 | 688.5 | 103.3 | 110.8 | 688.5 | 103.3 | 110.8 | Illumina HiSeq X Ten |
| PacBio | 1 | 8,944* | 1.7 | 14.8 | 15.9 | 1.7 | 14.8 | 15.9 | PacBio Sequel |
| Total | 17 | - | 3,581.9 | 398.8 | 427.8 | 3,339.5 | 350.5 | 376.0 | - |
| Assembly | # of contigs (w/o gaps) | N50 (contig) | # of gaps | total gap length | # of scaffolds | N50 (scaffold) | total length |
|---|---|---|---|---|---|---|---|
| 10X Chromium Genome linked-reads + PacBio reads | 34,366 | 161,334 | 23,539 | 43,071,641 | 10,823 | 20,385,050 | 999,596,594 |
| With added information from alignments against galGal5 | 21,892 | 174,303 | 19,355 | 36,492,537 | 2,538 | 81,982,712 | 932,102,785 |
| First version (all) | 305,507 | 28,461 | 57,682 | 17,858,319 | 247,825 | 3,678,261 | 937,605,096 |
| First version (>1k) | 65,900 | 30,310 | 56,401 | 17,810,012 | 9,499 | 3,851,064 | 898,141,602 |